RBF Neural Networks
Radial basis functions aren't just for meshfree PDE methods — they make excellent neural network activation functions. This page covers the theory behind RBF networks and walks through a hands-on comparison with a standard MLP using Lux.jl.
Why RBF Activations?
Local vs Global Activations
Standard activations like ReLU, tanh, and sigmoid are global: they partition the entire input space with hyperplanes, so every neuron responds to every input. Radial basis functions are local: each neuron is centered at a point in input space and responds most strongly to inputs near that center, decaying smoothly with distance.
This locality is particularly valuable for physics-informed neural networks (PINNs). Shouwei Li et al. (2024) introduced PIRBN (Physics-Informed Radial Basis Networks) and showed that PINNs naturally drift toward local approximation during training — RBF activations align with this tendency rather than fighting it, leading to faster convergence and better accuracy on PDEs.
The KAN–RBF Equivalence
Kolmogorov–Arnold Networks (KANs) attracted significant attention for placing learnable activation functions on edges rather than nodes. Li (2024) proved that KANs with RBF activations are mathematically equivalent to RBF networks, and that the simpler RBF formulation is faster in practice:
| Model | Test RMSE (function approx.) | Training time |
|---|---|---|
| Original KAN (B-spline) | Baseline | Baseline |
| FastKAN (Gaussian RBF) | Comparable | 3.3× faster |
| RBF-KAN (equivalence) | Comparable | Faster than KAN |
More recently, Putri et al. (2026) proved universal approximation for Free-RBF-KAN architectures with learnable centers, widths, and weights — confirming that RBF networks have the same expressive power as KANs without the implementation complexity.
Advantages for Scientific Computing
RBF activations inherit mathematical properties that standard activations lack:
| Property | RBF layer | Dense + ReLU/tanh |
|---|---|---|
| Smoothness | ||
| Locality | Each neuron covers a region | Every neuron responds globally |
| Interpolation guarantee | Exact with enough centers | No guarantee |
| Polynomial reproduction | Built-in via augmentation | Learned only |
| Known convergence rates | Yes (scattered data theory) | No |
| Interpretable parameters | Centers, shape params | Opaque weights |
Where RBF Layers Excel
RBF activations show the largest gains in scientific and geometric computing tasks:
| Application | Key result | Reference |
|---|---|---|
| PDE solving (PINNs) | 1000× better | RBF-PINN (Wang et al., 2024) |
| Physics-informed networks | Faster convergence on Poisson, Helmholtz, Burgers | PIRBN (Li et al., 2024) |
| Neural radiance fields | 10–100× fewer parameters for same quality | NeuRBF (Chen et al., 2023) |
| Neural fields (geometry) | Compact representation of SDFs and textures | NeuRBF (Chen et al., 2023) |
| Function approximation | Matches KAN accuracy, simpler architecture | FastKAN / RBF-KAN (Li, 2024) |
Learnable Shape Parameters
The shape parameter RBFLayer makes
Internally, the layer stores an unconstrained parameter
Initialization matters. Starting with
Training an RBF Network
This section trains an RBF network and a standard MLP on the same 1-D regression problem so you can compare convergence, accuracy, and interpretability.
Setup
using RadialBasisFunctions
using Lux, Optimisers, DifferentiationInterface, Mooncake
using Random, Statistics
using CairoMakie
const RBFLayer = Base.get_extension(
RadialBasisFunctions, :RadialBasisFunctionsLuxCoreExt
).RBFLayer
rng = Random.MersenneTwister(0)
# Target function with low- and high-frequency components
f(x) = sin(3x) + 0.3f0 * cos(7x)
# Training data: 50 points on [-1, 1]
n_train = 50
x_train = collect(Float32, range(-1, 1; length=n_train))
y_train = f.(x_train)
# Dense evaluation grid for plotting
x_plot = collect(Float32, range(-1, 1; length=300))
y_true = f.(x_plot)Model Definitions
Both models use roughly the same number of parameters so the comparison is fair.
# RBF network: 20 Gaussian centers
rbf_model = Chain(RBFLayer(1, 20, 1; basis_type=Gaussian))
# MLP: single hidden layer with relu activation
mlp_model = Chain(Dense(1 => 20, relu), Dense(20 => 1))
# Initialize
ps_rbf, st_rbf = Lux.setup(rng, rbf_model)
ps_mlp, st_mlp = Lux.setup(rng, mlp_model)
println("RBF parameters: ", Lux.parameterlength(rbf_model))
println("MLP parameters: ", Lux.parameterlength(mlp_model))RBF parameters: 61
MLP parameters: 61Training
A shared training loop keeps things comparable. Both models are trained with Adam for 1000 epochs on MSE loss.
function train(model, ps, st; lr=0.01f0, epochs=1000)
ps_flat, restructure = Optimisers.destructure(ps)
opt_state = Optimisers.setup(Adam(lr), ps_flat)
X = reshape(x_train, 1, :)
Y = reshape(y_train, 1, :)
backend = AutoMooncake(; config=nothing)
loss_fn(p) = mean((first(model(X, restructure(p), st)) .- Y) .^ 2)
losses = Float32[]
for epoch in 1:epochs
val, grads = DifferentiationInterface.value_and_gradient(loss_fn, backend, ps_flat)
push!(losses, val)
opt_state, ps_flat = Optimisers.update(opt_state, ps_flat, grads)
end
return restructure(ps_flat), losses
end
ps_rbf_trained, losses_rbf = train(rbf_model, ps_rbf, st_rbf)
ps_mlp_trained, losses_mlp = train(mlp_model, ps_mlp, st_mlp)
println("RBF final MSE: ", round(losses_rbf[end]; sigdigits=3))
println("MLP final MSE: ", round(losses_mlp[end]; sigdigits=3))RBF final MSE: 0.000338
MLP final MSE: 0.000911Loss Curves
fig = Figure(; size=(600, 350))
ax = Makie.Axis(fig[1, 1];
xlabel="Epoch", ylabel="MSE (log scale)",
yscale=log10, title="Training convergence")
lines!(ax, losses_rbf; label="RBF", linewidth=2)
lines!(ax, losses_mlp; label="MLP", linewidth=2, linestyle=:dash)
axislegend(ax; position=:rt)
fig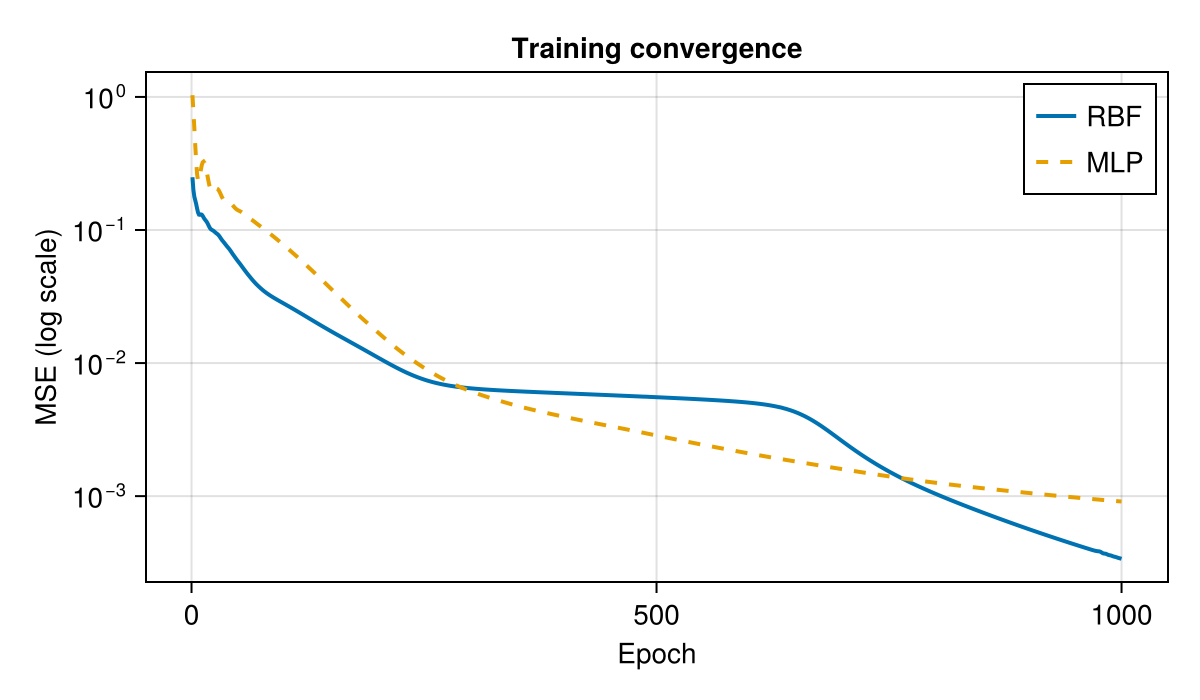
Final Fit
X_plot = reshape(x_plot, 1, :)
y_rbf, _ = rbf_model(X_plot, ps_rbf_trained, st_rbf)
y_mlp, _ = mlp_model(X_plot, ps_mlp_trained, st_mlp)
fig = Figure(; size=(700, 400))
ax = Makie.Axis(fig[1, 1]; xlabel="x", ylabel="f(x)", title="Learned fits after 1000 epochs")
lines!(ax, x_plot, y_true; label="Target", color=:black, linewidth=2)
lines!(ax, x_plot, vec(y_rbf); label="RBF", linewidth=2)
lines!(ax, x_plot, vec(y_mlp); label="MLP", linewidth=2, linestyle=:dash)
scatter!(ax, x_train, y_train; label="Training data", markersize=5, color=:gray)
axislegend(ax; position=:lb)
fig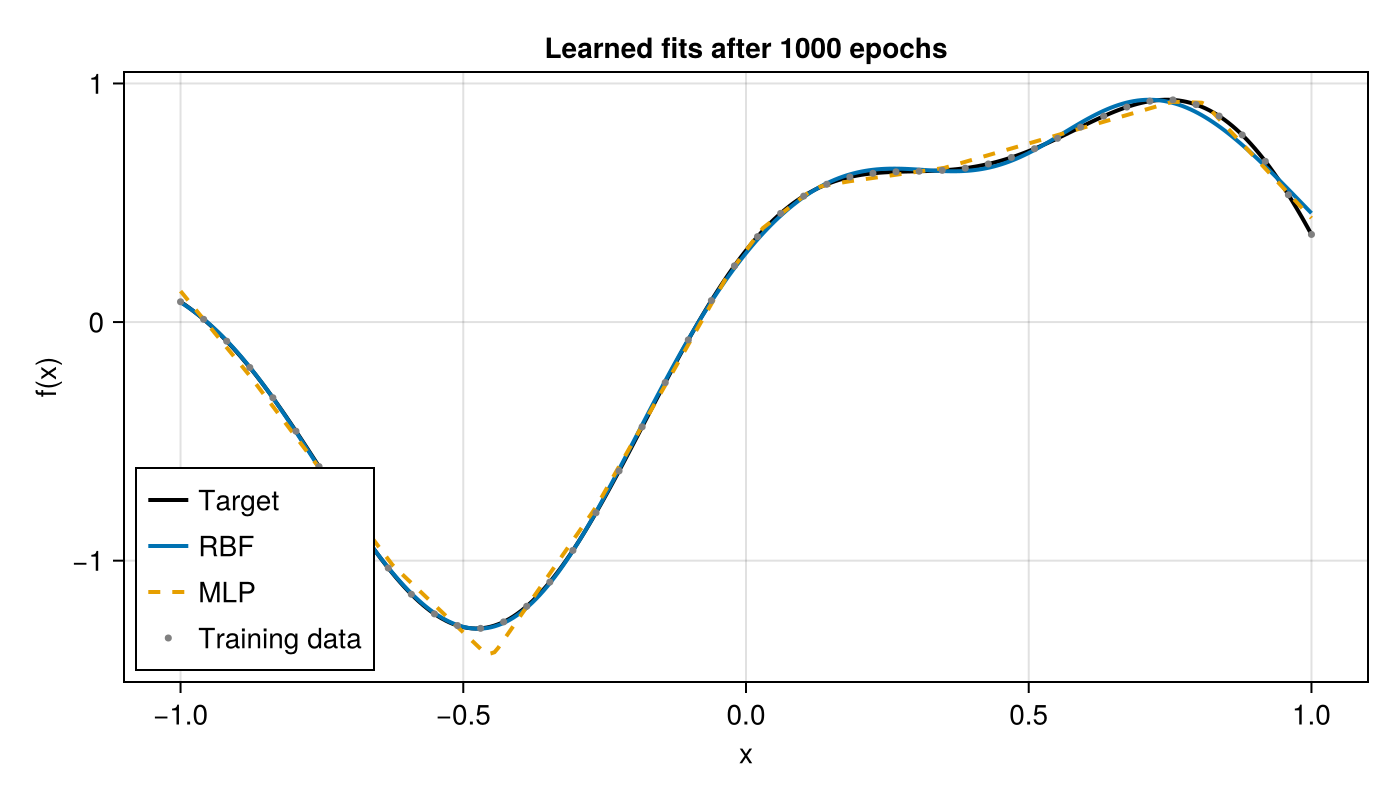
Learned Centers
A unique advantage of RBFLayer is that the centers are interpretable — each one anchors a basis function at a specific location in the input space.
centers = vec(ps_rbf_trained.layer_1.centers)
fig = Figure(; size=(700, 350))
ax = Makie.Axis(fig[1, 1]; xlabel="x", ylabel="f(x)", title="Learned RBF center locations")
lines!(ax, x_plot, y_true; color=:black, linewidth=2, label="Target")
vlines!(ax, centers; color=(:red, 0.5), linewidth=1.5, label="Centers")
axislegend(ax; position=:lb)
fig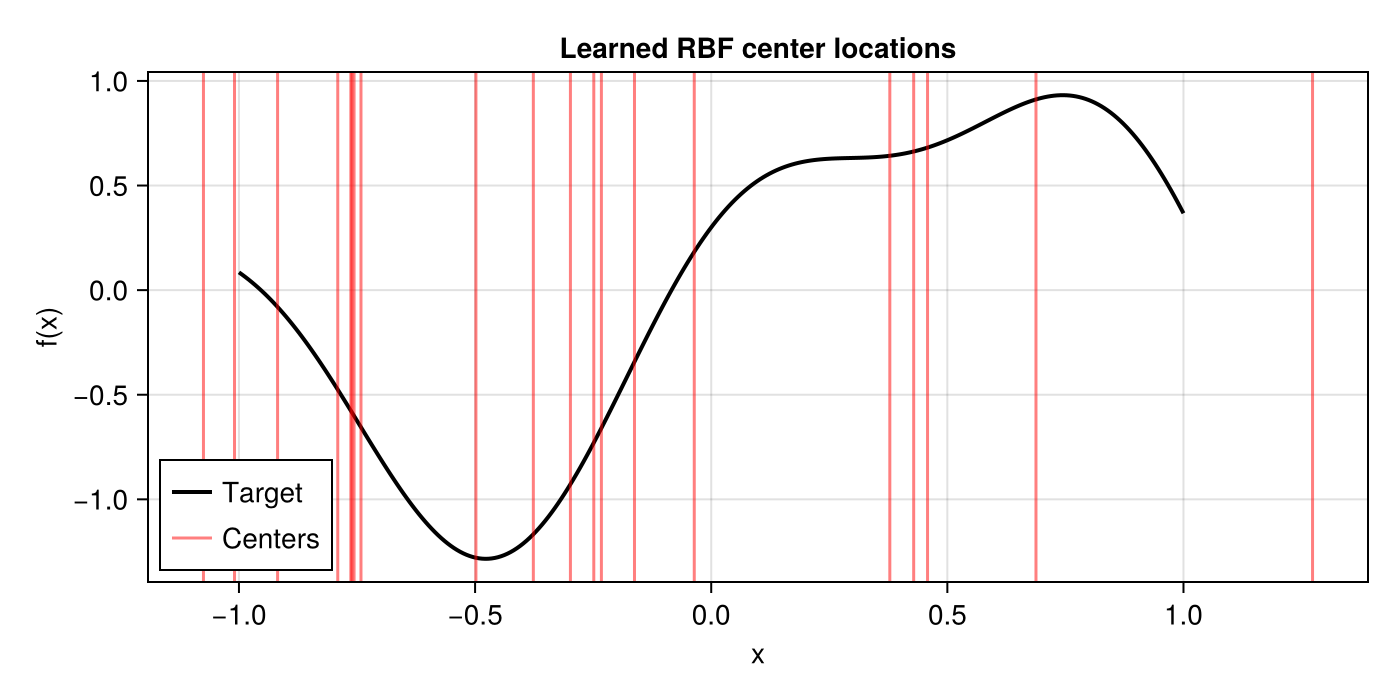
When to Use Standard Activations Instead
RBF layers are not universally superior. Prefer standard Dense + activation when:
High-dimensional, non-spatial inputs — Euclidean distance becomes less meaningful beyond ~10–20 dimensions (curse of dimensionality). Tabular data with mixed categorical/numerical features is better served by ReLU networks.
Deep architectures — Stacking many RBF layers is less studied than deep ReLU/transformer networks. For tasks requiring depth (NLP, large-scale vision), standard architectures have more mature tooling and theory.
Massive scale — When training on millions of samples with thousands of features, the per-neuron distance computation in RBF layers adds overhead compared to a simple matrix multiply + pointwise activation.
No geometric structure — If inputs have no notion of "closeness" (e.g., one-hot encoded categories, graph node IDs), locality provides no benefit.
References
S. Li, Y. Liu, & L. Liu, "PIRBN: Physics-Informed Radial Basis Networks," arXiv:2404.01445, 2024. Link
Z. Li, "Kolmogorov–Arnold Networks are Radial Basis Function Networks," arXiv:2405.06721, 2024. Link
J. Zhu, "FastKAN: Very Fast Kolmogorov-Arnold Networks," GitHub, 2024. Link
D. A. Putri, A. P. Tirtawardhana, & J. H. Yong, "Free-RBF-KAN," Engineering Applications of Artificial Intelligence, 2026. Link
Z. Wang, W. Xing, R. Kirby, & S. Zhe, "RBF-PINN: Non-Fourier Positional Embedding in Physics-Informed Neural Networks," arXiv:2402.08367, 2024. Link
Z. Chen, T. Li, Z. Ding, C. Wang, H. Bao, & Z. Chen, "NeuRBF: A Neural Fields Representation with Adaptive Radial Basis Functions," ICCV, 2023. Link